The results showed a general trend of sequence conservation as gene importance in function increased. This trend was somewhat similar when comparing to other genera.
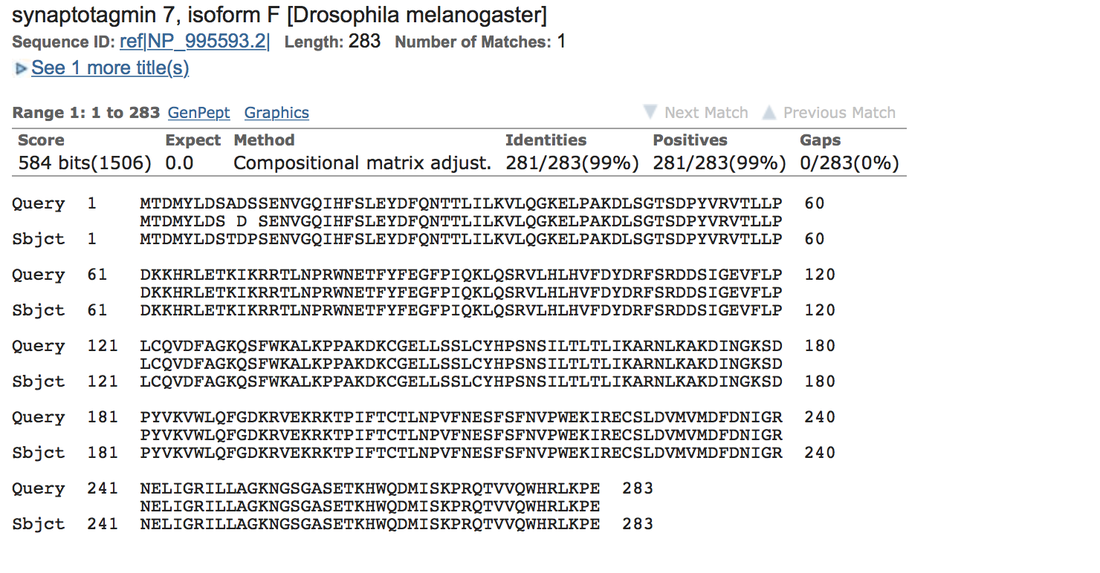
The Syt7 gene group in Drosophila elegans had the highest similarity and identity to Drosophila melanogaster. This may have been due to its function. Syt7 regulates synaptic transmission in the brain by transporting calcium ions, which causes stability. Its importance in the brain is the most likely reason the gene is conserved highly in Drosophila elegans and conserved relatively highly in other genera (HGNC, 2015).
The Syt7 gene group in Drosophila elegans had the highest similarity and identity to Drosophila melanogaster. This may have been due to its function. Syt7 regulates synaptic transmission in the brain by transporting calcium ions, which causes stability. Its importance in the brain is the most likely reason the gene is conserved highly in Drosophila elegans and conserved relatively highly in other genera (HGNC, 2015).
The next highly conserved gene is NfI. NfI is a protein that helps in hindering or expressing certain genes. This protein binds to genes making them inactive (Murtagh, J., Martin, F., & Gronostajski, R. M. 2003). This function’s importance may lead to high conservation.
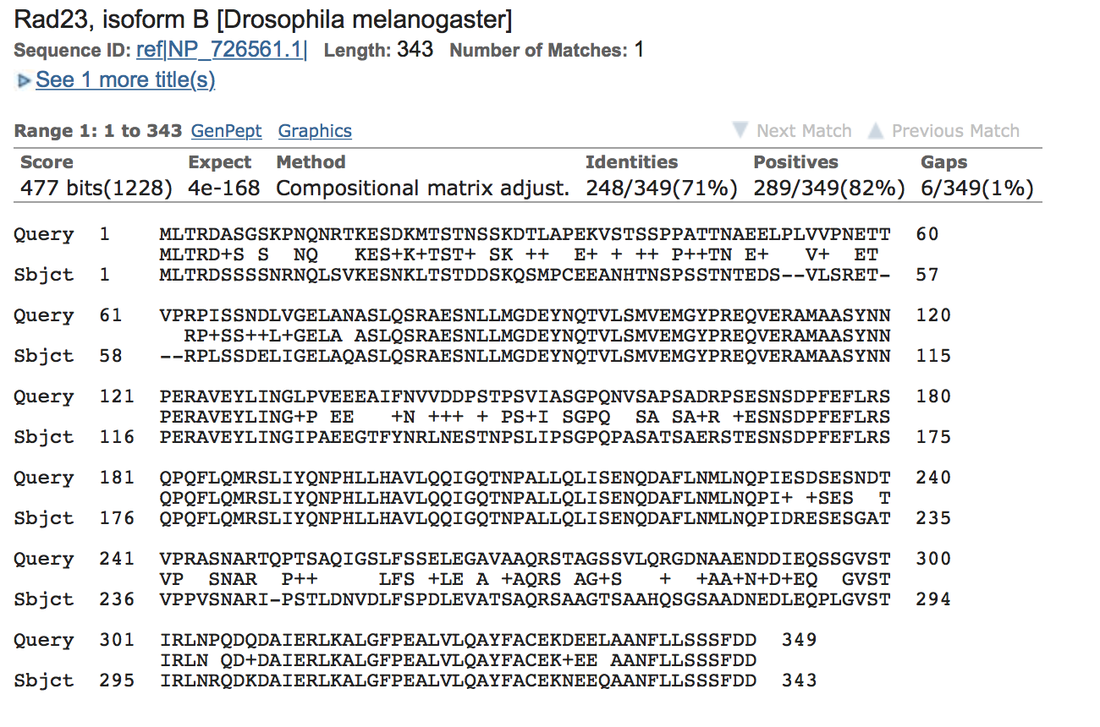
Rad23 is conserved more highly in other genera than within the same genus. Rad23 is involved in Nuclear Excision Repair, which is the recognition and repair of damaged DNA. (Wade, S., & Auble, D. 2010, August 1). This gene only protects some proteins from degradation. This gene is needed fairly often and is important for body function, which may lead to its high conservation.
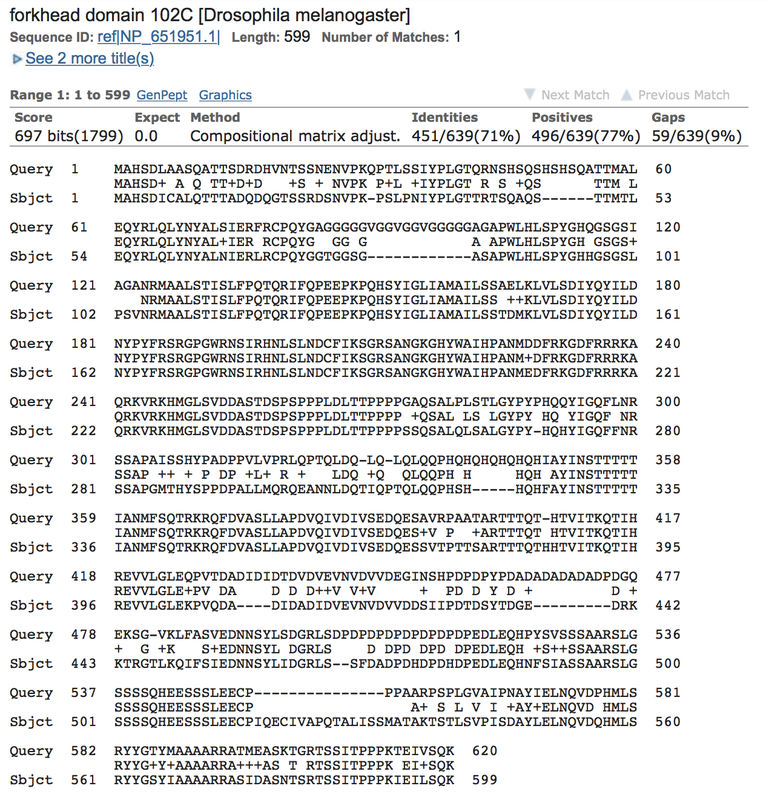
fd102c is the lowest conserved gene and this may be due to more reasons than just to function alone. The function of fd102c is for sequence specific DNA binding, transcription factor binding, and general DNA binding (Dos Santos, G., Schroeder, A. J., Goodman, J. L., Strelets, V. B., Crosby, M. A., Thurmond, J., FlyBase Consortium, T. F. 2014, November 14). Most of these functions were inferred from other similar genes, so more data is required to discern its true function. Based off of its function, it is not necessarily needed as much as other genes, which might be why it is minimally conserved. Another reason is that there is only one protein coding exon within the gene. There are no breaks in the sequence, so it is more prone to being altered, which may affect conservation.
CG11148 is the second least highly conserved gene. The function of the gene is unknown and has not been researched. So, in order to find a possible surrogate, the Raptor X prediction software was used These results gave 3 different templates for the gene from the two unique isoforms that CG11148 has. Each template has a different function. 4hpq:C predicts a protein that is involved in the breakdown of cellular structures. 3fma is a protein that binds and suppresses a gene in pre-mRNA process. 4h5y is a protein that binds to Rab proteins to help with their function. All of these templates give contradictory functions, but the best indicator is 4hpq:C due several data trends.
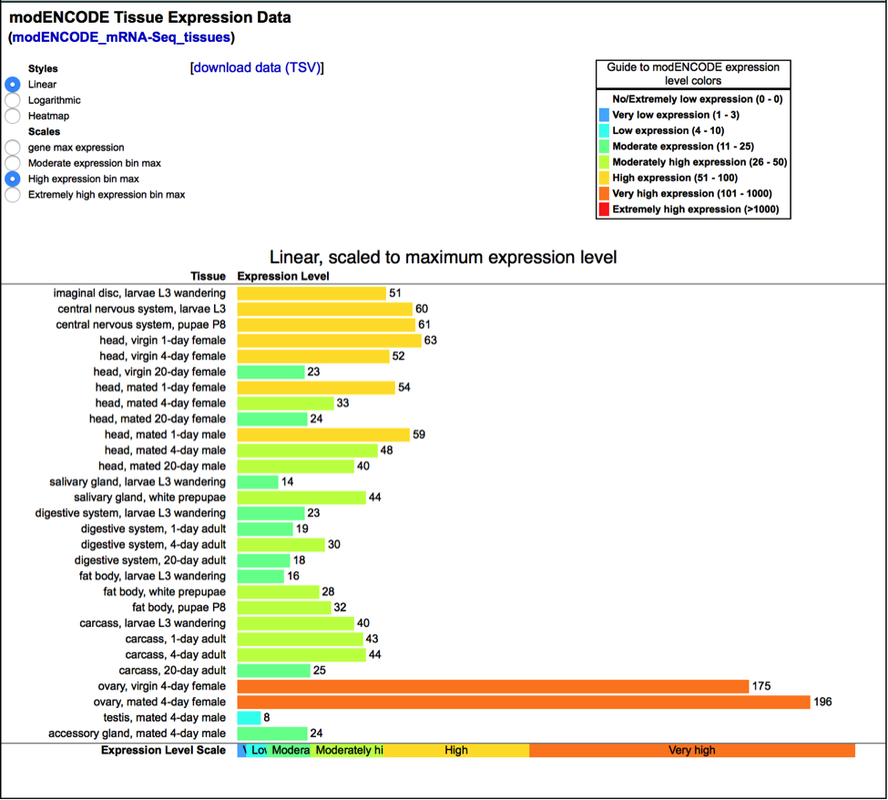
Within females, this gene is expressed very highly especially in the ovaries. In the head, after mating, this gene decreases in the head after several days (Dos Santos, G., Schroeder, A. J., Goodman, J. L., Strelets, V. B., Crosby, M. A., Thurmond, J., FlyBase Consortium, T. F. 2014, November 14). It was hypothesized that the protein derived from CG11148 was indeed a cell destruction protein, and it may be used to destroy and remove the embryo in females at birth. This is only a hypothesis so further testing is required. This function is somewhat important, which may be the reason for its lower sequence conservation in comparison to the other genes.